Gac Is a Transcriptional Repressor of the Lyme Disease Spirochete's OspC Virulence-Associated Surface Protein | Journal of Bacteriology

Structural basis of cyclic oligoadenylate binding to the transcription factor Csa3 outlines cross talk between type III and type I CRISPR systems - ScienceDirect

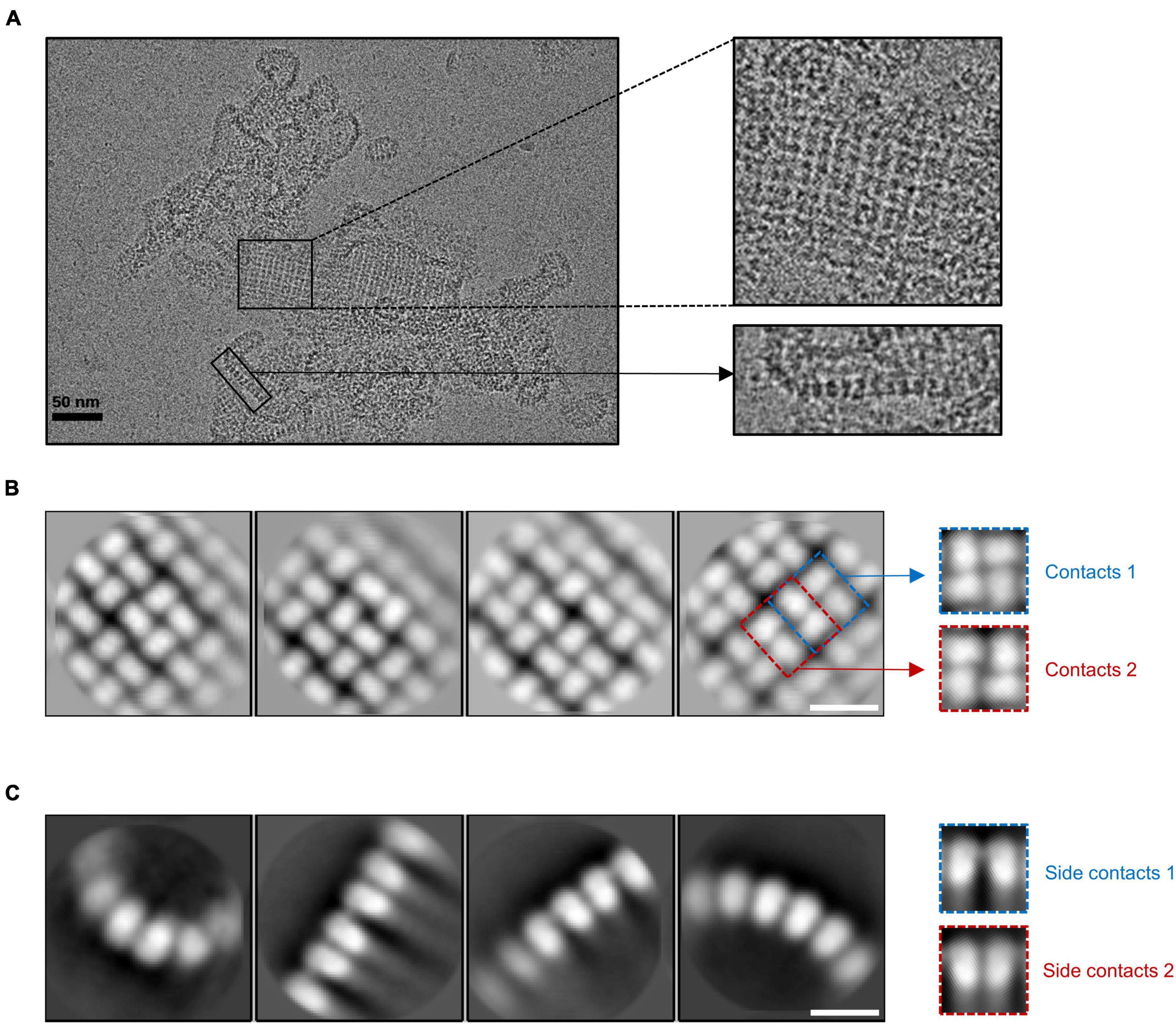
Biomolecules | Free Full-Text | Cyclic Tetra-Adenylate (cA4) Recognition by Csa3; Implications for an Integrated Class 1 CRISPR-Cas Immune Response in Saccharolobus solfataricus

IJMS | Free Full-Text | A Leucyl-tRNA Synthetase Urzyme: Authenticity of tRNA Synthetase Catalytic Activities and Promiscuous Phosphorylation of Leucyl-5′AMP

A Dimer Interface Mutation in Glyceraldehyde-3-Phosphate Dehydrogenase Regulates Its Binding to AU-rich RNA* - Journal of Biological Chemistry
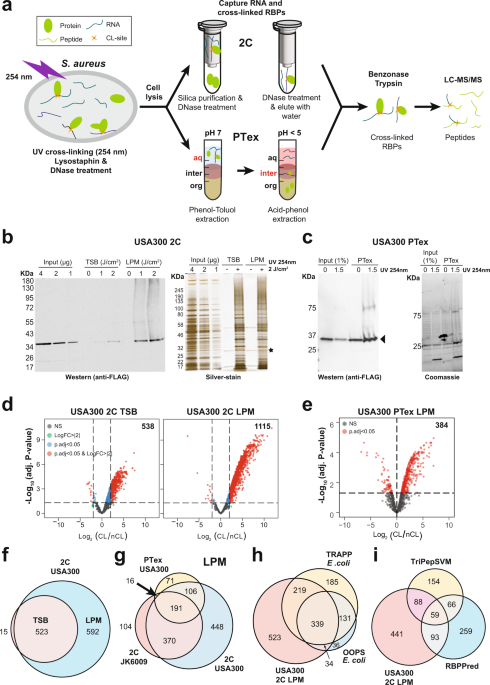
The RNA-bound proteome of MRSA reveals post-transcriptional roles for helix-turn-helix DNA-binding and Rossmann-fold proteins | Nature Communications
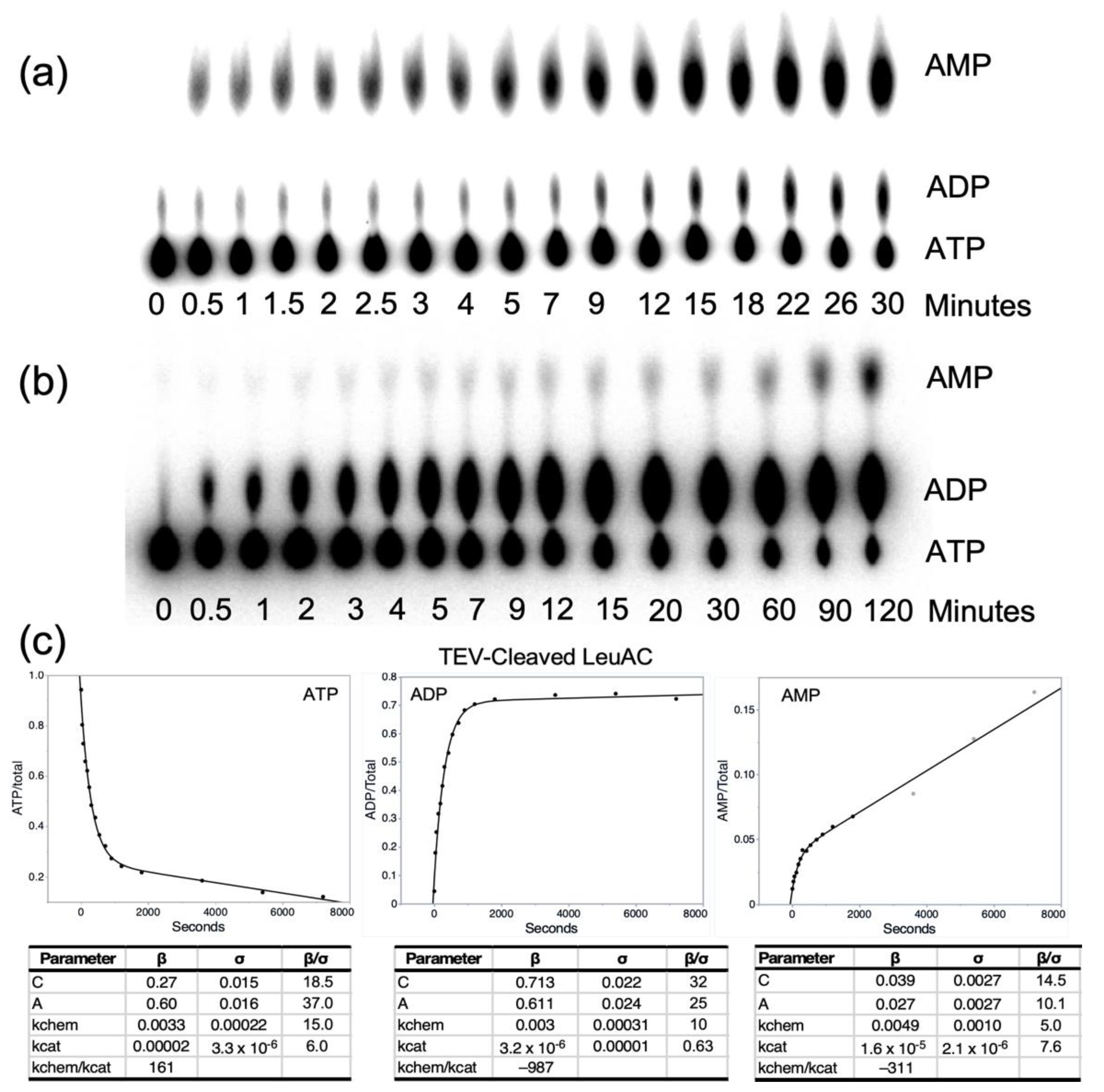
IJMS | Free Full-Text | A Leucyl-tRNA Synthetase Urzyme: Authenticity of tRNA Synthetase Catalytic Activities and Promiscuous Phosphorylation of Leucyl-5′AMP

Periodontitis Risk Variants at SIGLEC5 Impair ERG and MAFB Binding - R. Mueller, A. Chopra, H. Dommisch, A.S. Schaefer, 2022
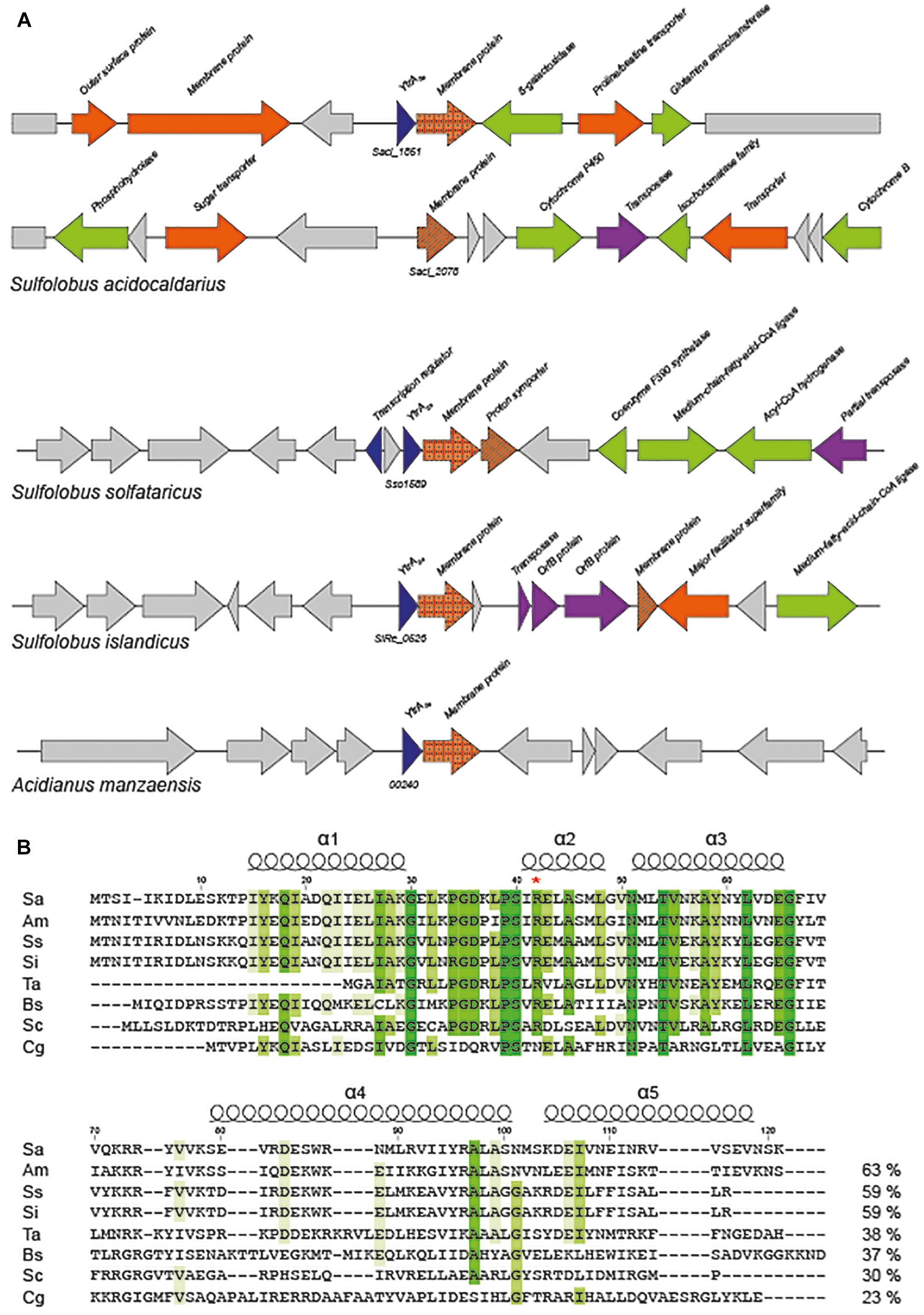
Frontiers | YtrASa, a GntR-Family Transcription Factor, Represses Two Genetic Loci Encoding Membrane Proteins in Sulfolobus acidocaldarius

Characterization of DNA Processing Protein A (DprA) of the Radiation-Resistant Bacterium Deinococcus radiodurans | Microbiology Spectrum









